FAIR 原則に基づく効率的なデータ管理をレイクハウスで実現

科学における発見とイノベーションの原動力となるデータ。そのデータの価値は、データ管理戦略によ��って左右されます。データ管理戦略は、データの品質、アクセスの容易さ、結果の再現性を確保するための重要な要素であり、信頼できる科学的なエビデンスを得るうえで欠かすことができません。
あらゆる分野の科学者が大規模なデータセットにアクセスできるようになったことで、管理や制御の煩雑さ、再現不能なデータワークフローをはじめとするビッグデータの問題が過去 10 年間で顕著になり、解決すべき課題となっています。
これを受けて、業界の専門家は、「価値あるデジタル資産の長期的な管理」を中核とした「適切なデータ管理とスチュワードシップ」のフレームワークを策定しました。このことは、2016 年に「Nature」誌の記事で初めて取り上げられています。このフレームワークは現在 FAIR 原則として認知されています。FAIR 原則は、デジタル資産の Findability(検索性)、Accessibility(アクセス性)、Interoperability(相互運用性)、Reusability(再利用性)が柱となっています。FAIR 原則のフレームワークでは、マシンの実行可能性および、人の介入なしで、あるいは最小限の介入だけで、データの発見、アクセス、相互運用、再利用を可能にするコンピューティングシステムの能力を中心に、前述の課題の解決を図ります。
科学分野のワークフローにおいて特定の問題を扱う場合、詳細なデータ品質管理から高度な分析まで、ほぼ全てのケースで新規に考案された統計手法を使用する必要があります。したがって、データアーキテクチャを適切なデータガバナンスに対応できるように設計するには、高度なデータ分析ツールの開発と応用についてもサポートしなければなりません。このような特徴に対しては従来の 2 層構造データアーキテクチャによる対応には限界があります。また、最新の高度なデータ分析ユースケースもサポートされていません。そこで役立つのが、レイクハウスアーキテクチャです。
ここ数年、データウェアハウスとデータレイクのメリットを新たなデータプラットフォームアーキテクチャに統合するレイクハウスのパラダイムが、業界全体に普及しつつあります。次世代のエンタープライズ規模のデータアーキテクチャが登場するなかで、レイクハウスは従来の分析や機械学習のユースケースをサポートできる汎用性の高い構造であることが証明されています。Delta Lake は、データレイクのためのオープンソースデータ管理レイヤーであり、データレイクによる拡張性、柔軟性、コスト削減効果に加え、データウェアハウスにみられる一貫性とトランザクション性能を提供します。
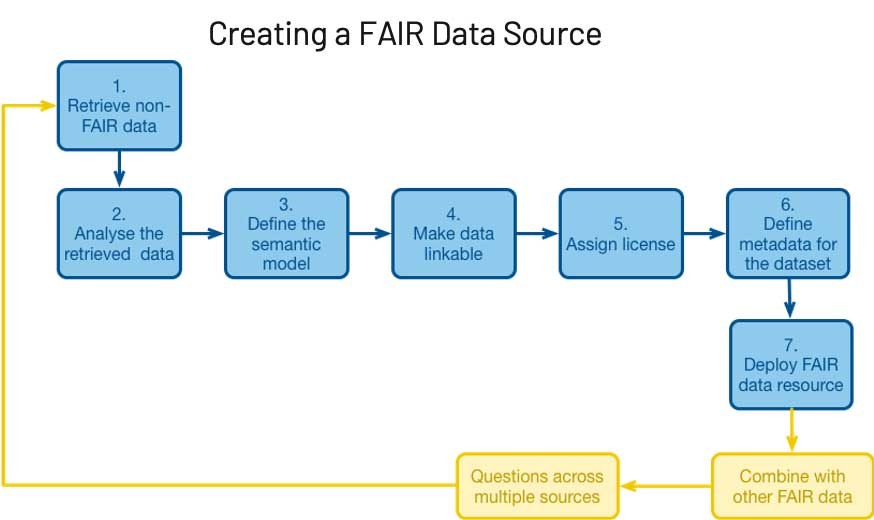
このブログ記事では、科学研究を進める組織のなかで、Delta Lake 上に構築されたレイクハウスが、FAIR 原則に基づいたデータシステムのアーキテクチャをどのように活用し、データ管理を実現しているかについて解説します。
レイクハウスの有用性は明白ですが、その実現については、長年にわたってデータチームの課題でした。例えば、データレイクです。データレイクほどアクセスが容易なシステムはありませんが、データ整理に大きなメリットをもたらした一方で、大きな混乱をもたらしました。また、クラウドにはさまざまなメリットがありますが、レイクハウスの実現をさらに難しくしています。クラウドを導入することで、ストレージコストが短期間のうちに下落し、場所や時間を問わないデータアクセスが実現しました。しかし、それがデータの拡散につながるという弊害が起こりました。このような状況が進むなかでは、高い理想を掲げたスチュワードシップの FAIR 原則は、優先順位が下がってしまいます。
クラウドの管理が十分でなければ、必ず弊害が生じます。コストの大幅な増加、利用率の低下、ガバナンスの欠如によるリスクへの対応が難しくなります。これは、あらゆる細胞、被験者、試験に不確実性と変化がつきものである科学の世界では特に顕著です。手元のパソコンに問題がないのに、不確実な要素をさらに増やす新たなデータプラットフォームの導入を考える人はいません。このように、データが未整理のままであることがイノベーションを妨げています。FAIR 原則では、このデータの整理を再現可能なプロセスにすることを目標にしています。では、FAIR 原則を実際に適用するにはどうすればよいのでしょうか?
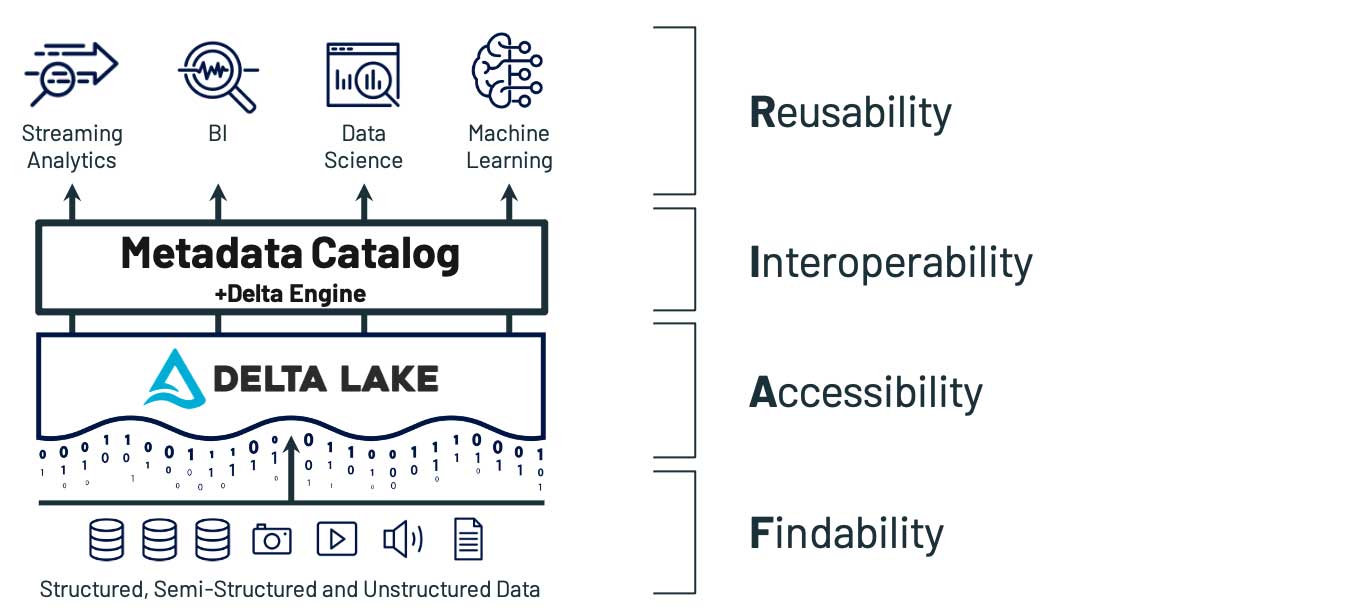
幸いなことに、近年のクラウドアーキテクチャの発展により、以前より容易に対応できるようになりました。Delta Lake 上に構築されたレイクハウスが FAIR の各原則にどのように対応しているか、具体的に見てみましょう。
Findability(検索性):自動化された反復可能な方法でデータを検索できる
実験、パイプライン、プロセスのいずれにおいても、最初に問題となるのがデータを見つけることです。これは、データの拡散で大きく妨げられます。ペタバイト単位のデータが何十もの独立したシステムに分散している状況では、企業内部の情報を把握していなければ対応は難しく、経験や知識のあるユーザーであっても、データ状況の把握に苦労するかもしれません。複数のシステムに分散しているデータの場所を一元化することは、データレイクの基本原則です。レイクハウスは、さらに FAIR の他の原則(Principles)もふまえて構築することで、この概念を拡張ますが、基本的な考え方は変わりません。適切に行えれば、�データを単一レイヤーに統合することで、他の全てのアーキテクチャの決定が容易になります。
FAIR 原則の Findability (検索性)に関する基準は、いくつかの副項目に分かれています。
- F1:(メタ)データにグローバルに一意で永続的な識別子(ID)が割り当てられている。
- F2:データがメタデータによって十分に記述されている。
- F3:メタデータが記述するデータの識別子(ID)が、メタデータに明確かつ明示的に含まれている。
- F4:(メタ)データが検索可能なリソースとして、登録もしくはインデックス化されている。
Delta Lake をベースとしたレイクハウスは、これら FAIR 原則の検索性の各項目に対応しています。例えば、Delta Lake では、スキーマなどの標準的な情報に加えて、バージョニング、スキーマの時系列での変化、ユーザーベースのリネージなどが、メタデータに含まれています。また、データとメタデータが同じ場所に格納されているため、データとそのデータを記述するメタデータとの関係が曖昧になることはありません。また、レイクハウスのベストプラクティスとして、一元化されたアクセス性の高いメタストアが用意されており、容易に検索できるようになっています。このように、レイクハウスの枠組みにおいては、高い水準の Findability(検索性) が実現されています。
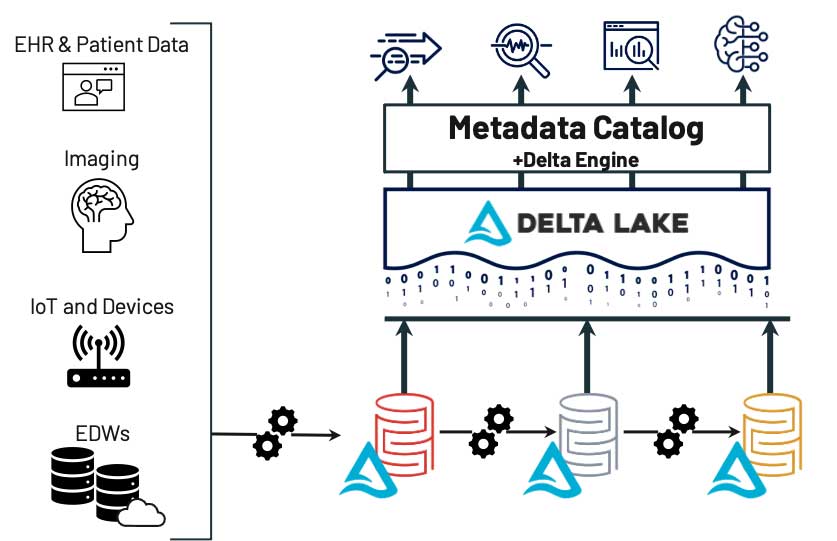
レイクハウスにより Findability(検索性) を実現している一例として、次のようなケースを考えてみましょう。
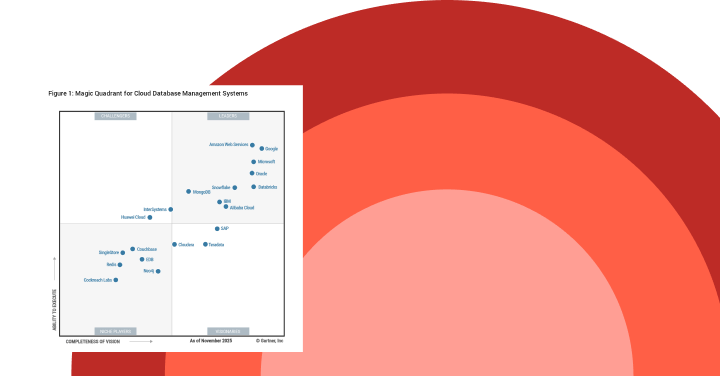
ガートナー®: Databricks、クラウドデータベースのリーダー
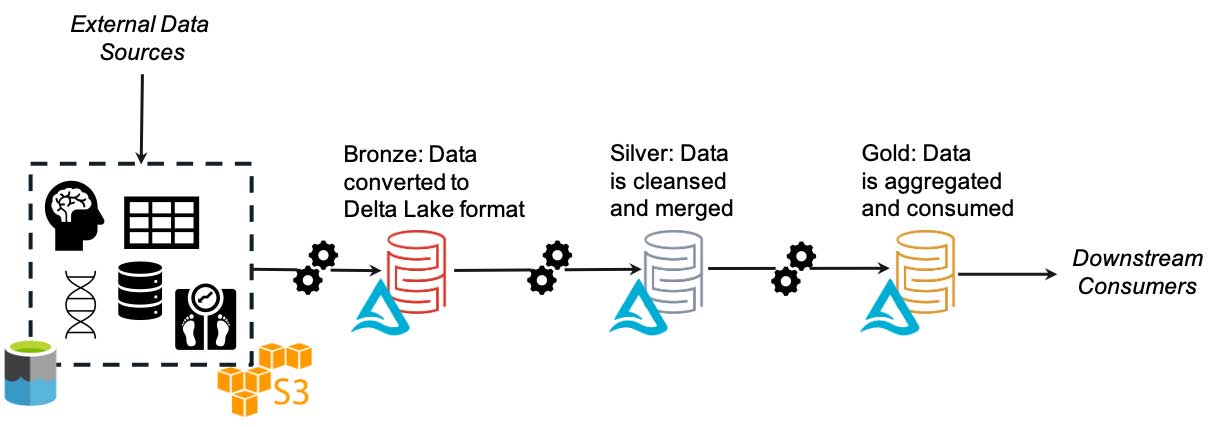
画像処理システム、オンプレミスやクラウドのデータウェアハウス、電子カルテ(EHR)システムなど、複数のシステムからデータが取得されているとします。ソースに関わらず、これらのデータは基礎となるデータレイク内のブロンズレイヤーに保存されます。その後、データは精製化のプロセスに自動で供給され、非特定化、正規化、フィルタリングなどが行われます。最終的に、データは高品質なデータのみを含むゴールドレイヤーに保存されます。ユーザー(または自動フィード)は 1 つの場所を検索するだけで、使用可能なデータの最新バージョンを見つけ�ることができます。データサイエンスや ML のプロセスにおいては、精製の程度の低いデータが必要になる場合がありますが、そのときにはシルバーレイヤーやブロンズレイヤーを活用できます。これらのプロセスでは、データの場所や各レイヤーの内容を確認できます。このような枠組みを用いることで、FAIR の他の全ての原則の実装と追跡が容易になります。
Accessibility(アクセス性):発見したデータにアクセスできる
FAIR 原則では、データへのアクセス性を「標準化された通信プロトコルを使って(中略)入手」し、「データが利用不可能となったとしてもアクセスできること」と定義しています。従来のデータレイクモデルが成立しなくなる原因は、この点でした。データレイクでは、任意の数のスキーマ、ファイルタイプ、フォーマット、データバージョンなどのほとんどが定義の段階で存在します。これは、Findability(検索性)には有用ですが、Accessibility(アクセス性)は困難になります。ある時点でデータレイクに存在していたものが、別の時点では変更、移動されていたり、完全に消えてしまったりということが起きます。これはデータレイクの主要な欠点の 1 つであり、レイクハウスに分岐する地点でもあります。
優れたレイクハウスのアーキテクチャには、基盤となるデータレイクとコンシューマの間に、Accessibility(アクセス��性)を促進するためのレイヤーが必要です。現在このようなレイヤーを提供するツールはいくつか存在しますが、最も広く使用されているのが Delta Lake です。Delta Lake には、ACID トランザクション、バッチとストリーミングの統合、クラウドに最適化された性能など、多くの利点があります。FAIR 原則との関連で特に重要なものは次の 2 点です。
1)Delta Lake は、Linux Foundation が管理するオープンソースのフォーマットであること。すなわち、独自仕様でなく、標準化されたプロトコルであり、マルチクラウドを念頭に設計されています。Delta Lake で生成されたデータは、ツールやベンダーを問わず、常にオープンなアクセスが可能です。
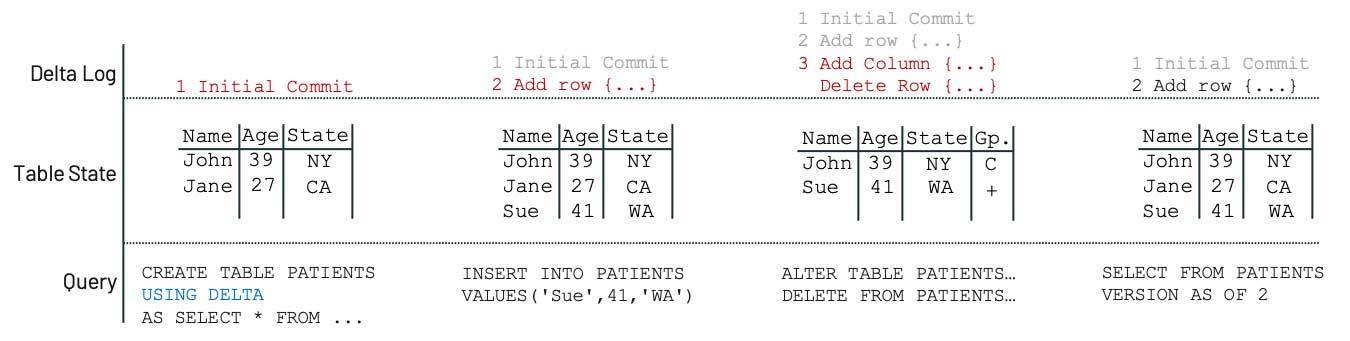
2)Delta Lake では、データとは別にトランザクションログが生成されること。このログは、再現性に必要なバージョニングなどのアクションに必要な情報を提供します。データが削除されても、メタデータの復元が可能となり、多くのケースでは、適切なバージョニングによりデータ自体の復元も可能となります。これは FAIR 原則である Accessibility(アクセス性)の要件を満たすうえで不可欠な要素です。時系列での安定性が保証されなければ、データは存在しないも同じとなってしまいます。
Delta Lake が Accessibility(アクセス性)を実現する例として、次のようなシナ��リオを考えてみましょう。患者情報のテーブルに新しいデータを追加した後、誤って意図しない変更を加えてしまったとします。
Delta Lake ではメタデータが永続化されており、変更に関するログも保持されているので、データを誤って削除した場合も、削除前の状態にアクセス可能です。テーブル全体が削除されたとしても同様です。これはシンプルな例ですが、Delta Lake 上に構築されたレイクハウスにより、データの安定性が確保され、Accessibility(アクセス性)が実現することがよくわかります。再現性が不可欠な組織にとっては特に有用であり、データチームの負担の軽減や、研究者の自由な探求とイノベーションが実現します。
さらに、Delta Lake では、セキュアなデータ共有のためのオープンプロトコルである Delta Sharing が提供されています。これにより、使用するコンピューティングプラットフォームに関わらず、管理しやすいオープンなフォーマットで、研究者が他の研究者や組織と研究データを直接共有することが容易になります。
Interoperability(相互運用性):データシステムが統合されている
データ��フォーマットは、近年、無数に存在します。かつては、CSV や表計算ソフト Excel 用など、使い慣れたフォーマットが必要な機能を全て提供していましたが、現在では BAM や SAM、HL7 など、医療分野に特化した何千ものフォーマットが存在します。また、DICOM 画像のような非構造化データ、Apache Parquet のようなビッグデータ標準、さらにはベンダー固有の独自フォーマットなど、数え切れないほどのフォーマットも存在します。これらのデータをデータレイクにまとめて投入すると、対処できないデータが混在する状況が生まれます。FAIR 原則を満たす実効性の高い相互運用システムでは、入力された全てのフォーマットが機械可読でなければなりません。しかし、HL7 で使用される膨大な種類のデータフォーマットは、機械可読が困難であり、一切判読できない場合もあります。
レイクハウスのパラダイムでは、Delta Lake を使ってこの問題に対処します。まず、データを元のフォーマットのまま保持し、履歴の記録やデータマイニングのために現状のコピーを保持します。次に、全てのデータを Delta フォーマットに変換し、下流のシステムが 1 つのフォーマットを認識するだけで機能するようにします。
レイクハウスでは、一元化された単一のメタデータカタログが用意されています。元データや変換済みデータの保存場所や保存方法にかかわらず、データの使用時・アクセス時に参照される場所が 1 つになります。さらに、厳格な PHI や HIPAA などのコンプライアンス要件を満たすデータに対しても単一の管理ポイントが存在することになり、データフローのガバナンスとコントロールが強化されます。
よくある質問に「これらの異なるフォーマットの全てを実際にどのように変換するか」というものがあります。下流のシステムで認識すべきフォーマットは Delta フォーマットのみですが、レイクハウスにおいて上流のデータを認識する仕組みが必要です。データブリックスでは、業界の専門家やパートナーと協力して、一般的に使用されるフォーマットを処理するためのソリューションを開発しました。ヘルスケアやライフサイエンスの分野のソリューションをいくつかご紹介します。
- GLOW:データブリックスとリジェネロンジェネティックセンター(Regeneron Genetic Center)の共同開発によるもので、一般的なゲノミクスフォーマットの取り込みと処理をスケーラブルかつ容易にし、より広範なデータと AI のエコシステムのなかでゲノミクスワークフローを容易に統合できるように設計されています。
- SMOLDER:HL7 データの取り込みと処理を目的とした Spark ベースのスケーラブルなフレームワークです。扱いが難しく変化しやすいフォーマットに対して使いやすいインタフェースを提供します。ネイティブリーダーとプラグインが提供されており、HL7 データが CSV ファイルと同様に利用できます。
Reusability(再利用性):複数のシナリオでデータの再利用ができる
Reusability(再利用性)については少し複雑です。既にレイクハウスアーキテクチャを構築している企業であっても、FAIR 原則に対処できない場合があります。これは、Reusability(再利用性)が技術的な問題以上のものだからです。ビジネスの中核を突き止め、難題をクリアする必要があります。ビジネスのサイロ化や部門間コラボレーションとチームワークを重視する企業風土の有無を考慮しなければなりません。研究開発部門と製造部門の責任者は、それぞれの部門でどのようにデータが使用されているかを把握できているでしょうか?レイクハウスがいかに優れていても、これらの問いに答えたり、その背景にある構造的な問題を解決したりすることはできません。しかし、それらの土台となる堅牢な基盤を提供することはできます。
レイクハウスが実現する価値は、データの取り込み、保存、バージョン管理、クリーニングなどの機能から生み出されるのではありません。ユースケースに関係なく、全てのデータの処理、アクセス、認識が可能な、単一の一元化プラットフォームが提供されることが重要なのです。データレイク、Delta Lake、Delta Engine、カタログなどの基盤となる部分は全て、これらのユースケースを実現するためのものです。優れたユースケースがなければ、どれだけ洗練されたアーキテクチャを持つデータプラットフォームであっても、価値を生み出すことはできません。
このブログでは、全てのデータのユースケースを網羅することはできませんでしたが、データブリックスのソリューションにより、科学データとコミュニティ標準の効果的な管理がどのようにして実現されているかを簡単にご紹介しました。レイクハウスによるソリューションについては、以下の入門編のリソースもご覧ください。
- ヘルスケアおよびライフサイエンスの Web ページ:業界におけるデータブリックスのソリューションやお客様事例をご紹介しています。
- バイオジェン(Biogen)における導入事例:バイオジェンでは、データブリックスのレイクハウスプラットフォームを新たな疾患の治療法の開発に活用しています。