Vectorized R I/O in Upcoming Apache Spark 3.0

R is one of the most popular computer languages in data science, specifically dedicated to statistical analysis with a number of extensions, such as RStudio addins and other R packages, for data processing and machine learning tasks. Moreover, it enables data scientists to easily visualize their data set.
By using SparkR in Apache SparkTM, R code can easily be scaled. To interactively run jobs, you can easily run the distributed computation by running an R shell.
When SparkR does not require interaction with the R process, the performance is virtually identical to other language APIs such as Scala, Java and Python. However, significant performance degradation happens when SparkR jobs interact with native R functions or data types.
Databricks Runtime introduced vectorization in SparkR to improve the performance of data I/O between Spark and R. We are excited to announce that using the R APIs from Apache Arrow 0.15.1, the vectorization is now available in the upcoming Apache Spark 3.0 with the substantial performance improvements.
This blog post outlines Spark and R interaction inside SparkR, the current native implementation and the vectorized implementation in SparkR with benchmark results.
Spark and R interaction
SparkR supports not only a rich set of ML and SQL-like APIs but also a set of APIs commonly used to directly interact with R code — for example, the seamless conversion of Spark DataFrame from/to R DataFrame, and the execution of R native functions on Spark DataFrame in a distributed manner.
In most cases, the performance is virtually consistent across other language APIs in Spark — for example, when user code relies on Spark UDFs and/or SQL APIs, the execution happens entirely inside the JVM with no performance penalty in I/O. See the cases below which take ~1 second similarly.
However, in cases where it requires to execute the R native function or convert it from/to R native types, the performance is hugely different as below.
createDataFrame()collect()dapply()dapplyCollect()gapply()gapplyCollect()
In short, createDataFrame() and collect() require to (de)serialize and convert the data from JVM from/to R driver side. For example, String in Java becomes character in R. For dapply() and gapply(), the conversion between JVM and R executors is required because it needs to (de)serialize both R native function and the data. In case of dapplyCollect() and gapplyCollect(), it requires the overhead at both driver and executors between JVM and R.
Native implementation
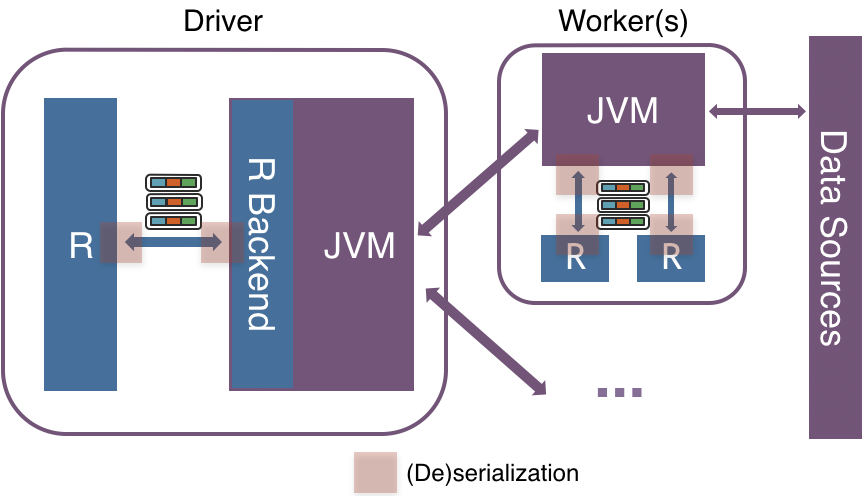
The computation on SparkR DataFrame gets distributed across all the nodes available on the Spark cluster. There’s no communication with the R processes above in driver or executor sides if it does not need to collect data as R data.frame or to execute R native functions. When it requires R data.frame or the execution of R native function, they communicate using sockets between JVM and R driver/executors.
It (de)serializes and transfers data row by row between JVM and R with an inefficient encoding format, which does not take the modern CPU design into account such as CPU pipelining.
Vectorized implementation
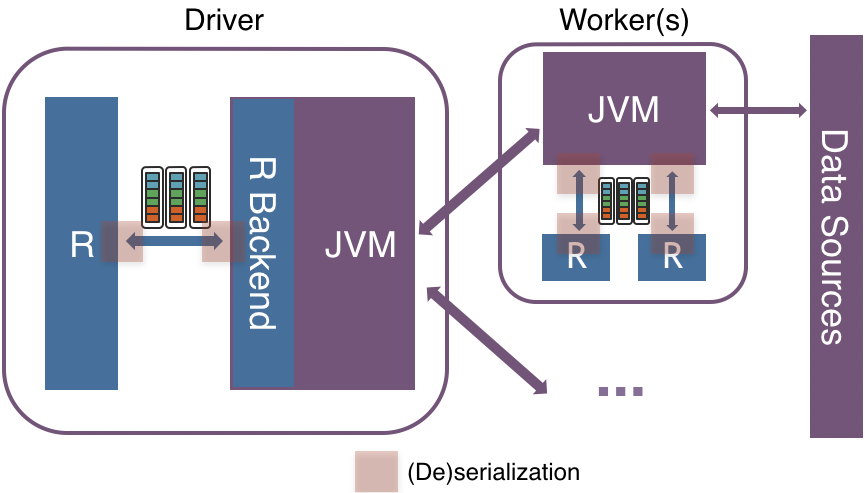
In Apache Spark 3.0, a new vectorized implementation is introduced in SparkR by leveraging Apache Arrow to exchange data directly between JVM and R driver/executors with minimal (de)serialization cost.
Instead of (de)serializing the data row by row using an inefficient format between JVM and R, the new implementation leverages Apache Arrow to allow pipelining and Single Instruction Multiple Data (SIMD) with an efficient columnar format.
The new vectorized SparkR APIs are not enabled by default but can be enabled by setting spark.sql.execution.arrow.sparkr.enabled to true in the upcoming Apache Spark 3.0. Note that vectorized dapplyCollect() and gapplyCollect() are not implemented yet. It is encouraged for users to use dapply() and gapply() instead.
Benchmark results
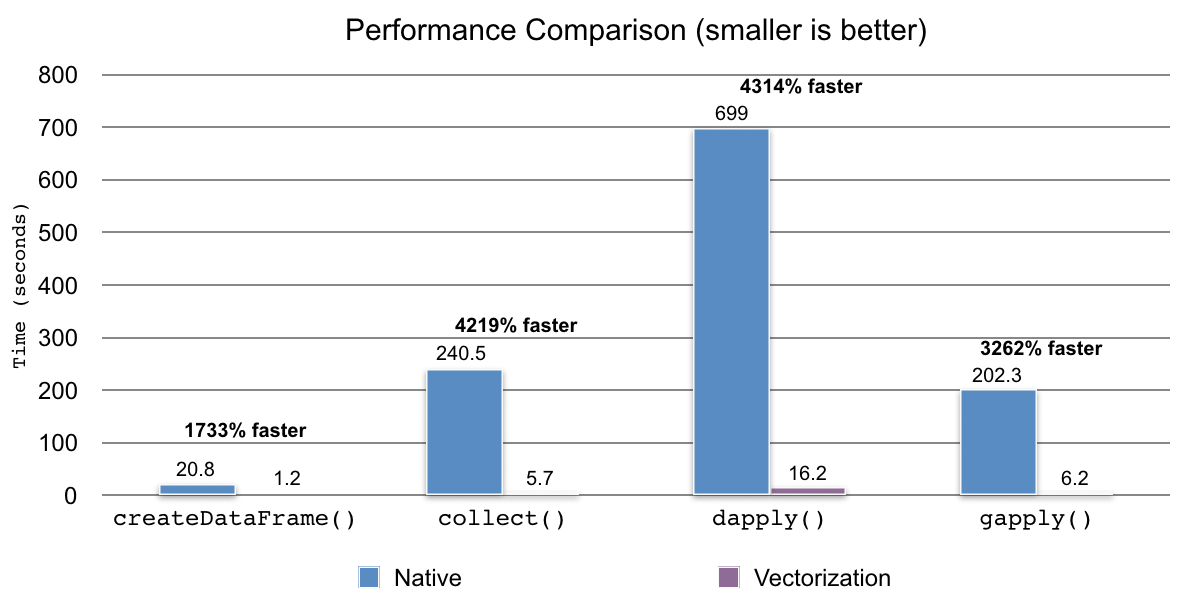
The benchmarks were performed with a simple data set of 500,000 records by executing the same code and comparing the total elapsed times when the vectorization is enabled and disabled. Our code, dataset and notebooks are available here on GitHub.
In case of collect() and createDataFrame() with R DataFrame, it became approximately 17x and 42x faster when the vectorization was enabled. For dapply() and gapply(), it was 43x and 33x faster than when the vectorization is disabled, respectively.
There was a performance improvement of up to 17x–43x when the optimization was enabled by spark.sql.execution.arrow.sparkr.enabled to true. The larger the data was, the higher performance expected. For details, see the benchmark performed previously for Databricks Runtime.
Conclusion
The upcoming Apache Spark 3.0, supports the vectorized APIs, dapply(), gapply(), collect() and createDataFrame() with R DataFrame by leveraging Apache Arrow. Enabling vectorization in SparkR improved the performance up to 43x faster, and more boost is expected when the size of data is larger.
As for future work, there is an ongoing issue in Apache Arrow, ARROW-4512. The communication between JVM and R is not fully in a streaming manner currently. It has to (de)serialize in batch because Arrow R API does not support this out of the box. In addition, dapplyCollect() and gapplyCollect() will be supported in Apache Spark 3.x releases. Users can work around via dapply() and collect(), and gapply() and collect() individually in the meantime.
Try out these new capabilities today on Databricks, through our DBR 7.0 Beta, which includes a preview of the upcoming Spark 3.0 release. Learn more about Spark 3.0 in our preview webinar.